Macromolecular Geometry
 Overview
Overview
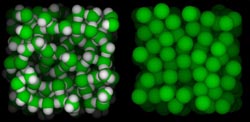
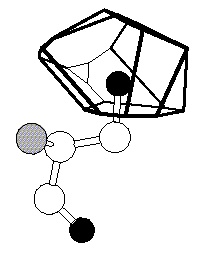
We are interested in the calculation of geometrical quantities
associated with macromolecular structures and their motions.
These include volumes, surfaces, axes, angles, and
distances. Volumes and other quantities related to packing
are of particular interest. For instance, one goal is to
characterize the special type of atom-to-atom packing that
occurs at interfaces, such as those between a protein and
water. This analysis is motivated by the work on protein
packing begun by Fred
Richards a quarter century ago.
 Information
Information
 Papers
Papers
- M Gerstein (1992). "A Resolution-Sensitive Procedure for
Comparing Protein Surfaces and its Application to the
Comparison of Antigen-Combining Sites," Acta Cryst.
A48: 271-276.
- [Abstract] [ftp directory with manuscript]
[Talk
Abstract]
- M Gerstein & R M Lynden-Bell (1993). "Simulation of
Water around a Model Protein Helix. 2. The Relative
Contributions of Packing, Hydrophobicity, and Hydrogen
Bonding," J. Phys. Chem. 97: 2991-2999.
- [Abstract]
- M Gerstein & R M Lynden-Bell (1993). "What is the
natural boundary for a protein in solution?" J. Mol.
Biol. 230: 641-650.
- M Gerstein, A M Lesk, E N Baker, B Anderson, G Norris
& C Chothia (1993). "Domain Closure in Lactoferrin: Two
Hinges produce a See-saw Motion between Alternative
Close-Packed Interfaces," J. Mol. Biol. 234:
357-372.
- [Medline
Abstract for 94047086] [Summary Graphic] [ftp directory with many
pictures]
- Y Harpaz, M Gerstein & C Chothia (1994). "Volume
Changes on Protein Folding," Structure 2:
641-649.
- [Medline
Abstract for 95006332]
- M Gerstein, J Tsai & M Levitt (1995). "The volume of
atoms on the protein surface: Calculated from simulation,
using Voronoi polyhedra," J. Mol. Biol. 249:
955-966.
- [ftp directory with
manuscript, talks, programs & data]
- M Suzuki & M Gerstein (1995). "The geometry of
alpha-helices binding to DNA," Proteins: Structure,
Function, Genetics 23: 525-535.
- J Tsai, M Gerstein & M Levitt (1996). "Keeping the
Shape but Changing the Charges: A Simulation Study of Urea
and its Iso-steric Analogues," J. Chem. Phys.
104: 9417-9430.
- M Gerstein & C Chothia (1996). "Packing at the
Protein-Water Interface" Proc. Natl. Acad. Sci. USA
93: 10167-10172.
- [PubMed
8816770]
 Source Code
Source Code
A library of C source code is available. It contains programs
for calculating volumes (using the Voronoi method), surfaces
contacts, and other parameters relevant to the protein surface.
It also contains code for doing a least-squares fit of two
structures and for calculating geometrical parameters such as
helix axes.
Also, there is:
Check out "beta" of version
2.0 of source code release.
 Standard
volumes
Standard
volumes
A file
giving statistics on the volumes for buried core atoms in a
database of 119 structures. These reference atomic volumes can
be compared to the volumes calculated with the volume program
(above) to see how well-packed a given atom is.
28 January 2003 / graham@bioinfo.mbb.yale.edu
1 May 1997 / Mark.Gerstein@Yale.edu
 Overview
Overview
 Overview
Overview Information
Information Papers
Papers Source Code
Source Code Standard
volumes
Standard
volumes Picture Gallery
Picture Gallery