(click picture for expanded view)

(click on picture to enlarge)
confirmed by a second investigator, and
expressed as a single digit numeric. A part of the bands were quantified using
PhosphorImager System (Molecular Dynamics).
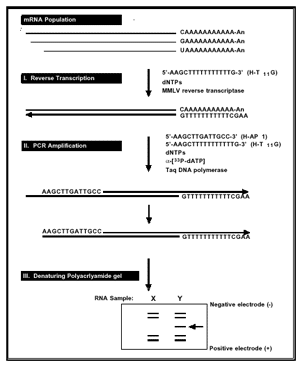
image obtained from www.genehunter.com