Comparer for protein structure rankings
Qian, J. et.al (2001) Nucleic Acids Research 29: 1750-64|
[First 30 records][Entire table] [Display values][Display ranks] Image size: [50][70][100][120] | U(all) untrimmed RMS | R(all) trimmed RMS | Q(all) P-value (seq.) | B(Gly,pdb40) composition percentage of Gly for pdb40d | B(His,pdb100) composition percentage of His for pdb100d | |
| Max in column | 25.2 | 10.3 | 9.8.e-01 | 18.1 | 8.7 | |
| Min in column | 0.14 | 0.06 | 1.51E-260 | 1.5 | 0.1 | |
| Average | 6.6 | 1.8 | 0.8 | 7.5 | 2.4 | |
| Non zero hits | 216 | 216 | 216 | 407 | 394 | |
| Ranked folds (click on arrows to re-rank) |  |  |  |  |  | |
 |  |  |  |  | ||
d2pgd_1:. (1.39) |  | 25.2 | 3.51 | 1 | 8.1 | 2.3 |
d1aj3__:. (1.39) |  | 24.72 | 2.78 | 7.5.e-01 | 5.9 | 4.4 |
d1poib_:. (1.39) |  | 20.52 | 10.3 | 1 | 9.2 | 2.3 |
d1gtqa_:. (1.39) |  | 19.33 | 10.01 | 1 | 4.7 | 4.5 |
d1a0p__:. (1.39) |  | 18.78 | 3.09 | 1 | 5.2 | 3.4 |
d1pysa_:. (1.39) |  | 17.49 | 6.95 | 9.14E-01 | 8.1 | 2.2 |
d1tfe__:. (1.39) |  | 16.61 | 8.71 | 1.11E-01 | 7.3 | 1.9 |
d1a8e__:. (1.39) | 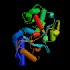 | 16.26 | 7.53 | 7.89E-01 | 6.9 | 1.2 |
d1pkp_1:. (1.39) |  | 15.66 | 5.34 | 1 | 10.1 | 1.7 |
d1dhx__:. (1.39) |  | 15.65 | 5.99 | 9.14E-01 | 6.7 | 2.3 |
d1puc__:. (1.39) |  | 15.19 | 0.41 | 1.80E-15 | 4.8 | 5.9 |
d1eur__:. (1.39) |  | 14.64 | 6.67 | 6.00E-01 | 9.5 | 2.1 |
d1nsya_:. (1.39) |  | 14.62 | 6.02 | 8.75E-01 | 7.5 | 2.7 |
d2lbd__:. (1.39) |  | 14.18 | 2.02 | 1.03E-07 | 5.0 | 3.5 |
d1prta_:. (1.39) |  | 13.58 | 5.69 | 9.8.e-01 | 8.8 | 2.3 |
d1rec__:. (1.39) |  | 13.45 | 3.35 | 4.12E-01 | 6.3 | 1.4 |
d1hmy__:. (1.39) |  | 13.45 | 4.02 | 9.71E-01 | 6.6 | 2.7 |
d1oroa_:. (1.39) |  | 13.41 | 4.88 | 7.88E-01 | 7.2 | 1.7 |
d2rig__:. (1.39) |  | 12.62 | 3.26 | 8.66E-01 | 3.6 | 2.4 |
d2lbp__:. (1.39) |  | 12.61 | 4.82 | 6.33E-01 | 8.4 | 1.5 |
d1ddf__:. (1.39) |  | 12.52 | 2.87 | 1 | 5.9 | 5.2 |
d1azo__:. (1.39) |  | 12.52 | 3.47 | 1 | 6.7 | 2.3 |
d2xat__:. (1.39) |  | 12.48 | 1.44 | 1 | 10.0 | 2.5 |
d1xsm__:. (1.39) |  | 12.18 | 4.4 | 8.88E-01 | 5.6 | 2.8 |
d1rfs__:. (1.39) |  | 11.62 | 1.99 | 6.05E-07 | 8.5 | 3.4 |
d1leha2:. (1.39) |  | 11.5 | 2.39 | 4.15E-01 | 9.9 | 1.4 |
d1neda_:. (1.39) |  | 11.24 | 2.07 | 7.93E-01 | 8.6 | 2.1 |
d1bak__:. (1.39) |  | 11.24 | 3.25 | 8.59E-01 | 5.7 | 2.4 |
d1bgp__:. (1.39) |  | 11.18 | 2.18 | 4.48E-01 | 7.6 | 2.3 |
d1alo_3:. (1.39) |  | 11.18 | 3.3 | 1 | 8.1 | 2.3 |
The images of protein structures were taken from www.rcsb.org
The classification of folds is from SCOP version 1.39
Questions, comments, and suggestions qian@csb.yale.edu